Source Control for Data Science - using Azure DevOps / VSTS with Jupyter Notebooks
So many of you will know about https://mybinder.org/ Binder is a awesome tool that allows you turn a Git repo into a collection of interactive Jupyter notebooks and it allows you to, open those notebooks in an executable environment, making your code immediately reproducible by anyone, anywhere.
Jupyter Notebooks in the cloud
Another great interactive tool for working with Notebooks is Microsoft Azure Notebook https://notebooks.azure.com
Integrate notebooks with Git
However in many cases Data Scientists want to work collaboratively and then publish, share both their notebooks and results. This is where using some form of source control is vital.
Here are a few suggestions and tips regarding how make process of notebooks PR reviews and overall co-existence with git more productive and easy.
1) Minimal requirement would be to manually always clear all outputs before pushing to repo.
2) This link talks about how to automate this: https://timstaley.co.uk/posts/making-git-and-jupyter-notebooks-play-nice/
3) Additional useful option would be to have "save hook" that automatically saves python-only script ver of the notebook (for future easier code reviews) as well as html ver (so it is possible to see the outputs), see ideas here https://www.svds.com/jupyter-notebook-best-practices-for-data-science/
So here is an example of this in practice
Setup Git Hooks
- Ensure you are running Git version
2.9or greater - Run the
Makefilewithmake
First of all you need to create a new Makefile in the route of your repo
init: git config core.hooksPath .githooks
Then need you need to create a .githooks folder within the root
with a pre-commit file
#!/bin/bash
IPYNB_FILES=`git diff --staged --name-only | awk "/.ipynb/"`echo "$IPYNB_FILES"
cd "notebooks"
function get_script_extension() { # find file_extension from notebook metadata json, get value from file_extension property echo $(awk '/"file_extension": ".*"/{ print $0 }' $1 | cut -d '"' -f4)}
if [[ -n "$IPYNB_FILES" ]]; then for ipynb_file in $IPYNB_FILES do file_path="${ipynb_file##*/}" file_no_extension="${file_path%.*}"
# if file exists and is not a checkpoint file if [[ -f "$file_path" ]] && [[ "$file_path" != *"checkpoint"* ]]; then echo "Creating scripts from notebook." jupyter nbconvert --to script "$file_path" # update scripts file in scripts directory extension=$(get_script_extension $file_path) mv -f "$file_no_extension$extension" "../scripts"
echo "Creating html from notebook." jupyter nbconvert --to html "$file_path" # update html file in scripts directory mv -f "$file_no_extension.html" "../html"
# clear notebook output echo "Clearing notebook output." jupyter nbconvert --ClearOutputPreprocessor.enabled=True --inplace "$file_no_extension.ipynb" fi donefi
# return to root directorycd ..
# add the newly created files to the current commitgit add -A
Within your repo you simply have the following three folders
Html – where a full rendered notebook is stored
Scripts – Where you R Scripts are stored
Notebooks – where you ipynb are stored
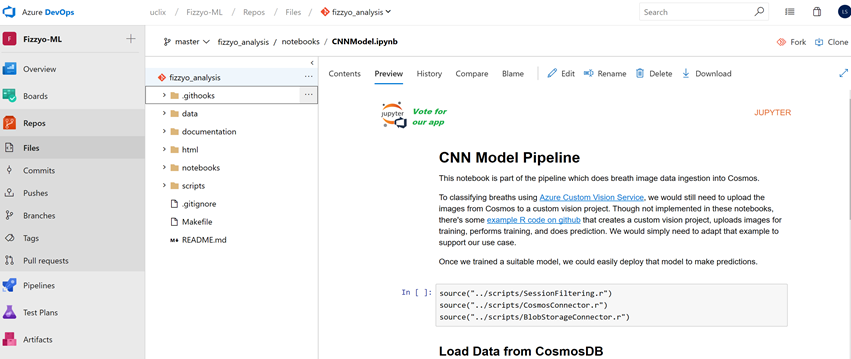
Enable Jupyter Notebook support in Azure DevOps
So this is the perfect layout for storing your experiments within GitHub or Azure Devops but one of the most frustrating things was viewing your output within Azure Devops/VSTS see https://developercommunity.visualstudio.com/content/idea/365561/add-support-for-viewing-jupyter-ipython-notebooks.html
The Microsoft team have developed a FREE VSTS Marketplace extension that will allow data scientists to use VSTS for source control and allow them to preview their Jupyter Notebooks from within VSTS (or TFS)
You simply install the extension to your Azure DevOps account https://marketplace.visualstudio.com/items?itemName=ms-air-aiagility.ipynb-renderer
Choose any Jupyter Notebook (.ipynb) file in your repo. The preview pane will show the rendered content of Jupyter Notebook.
Conclusion
As you can see this makes using Azure DevOps ideal for data Scientist to collaborate and share resources and code across and experiment whilst using the githooks to ensure that they always have evidence based outcomes of each iteration and with the extension the notebooks can shared and discussed.