Using Concurrency runtime, and Visual Studio to build a financial application
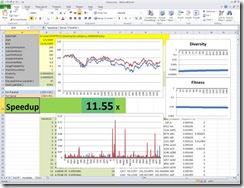
Figure 1: A screen shoot on a 16 core Windows 7
This blog is about an end-to-end scenario for the Concurrency Runtime and Visual Studio 2010, involving building a financial application, and measuring and tuning its performance.
The user scenario for this application is that an investor is looking for a subset of instruments in an index like the S&P500 that best approximates the overall index’s behavior. Due to an almost infinite number of permutations, including which subsets of instruments to consider as well as how much investment should be made in each instrument, it is near impossible to have a closed form solution for this problem. A heuristic is needed, and in this case the developer implementing a solution to this scenario uses a form of genetic algorithm called Differential Evolution.
We will take the S & P 500 as an example, and to apply differential evolution to our problem we will make these substitutions:
- Complete S & P 500 = Chromosomes
- Max Contributors = non zero chromosomes
- Num. of Portfolios = num. of individuals
- Num. of iterations = num. of generations
Differential Evolution creates a random population, then continues to evolve that population until a certain number of iterations (generations) is reached or until a satisfactory result is found. An evolution consists of four steps executed on each individual.
- Mutation: Selecting 3 individuals (donors) to pick their chromosomes.
- Recombination: randomly copy chromosomes from the one individual to the newly generated one
- Reparation: re-apply the rules back after mutation, and recombination
- Selection: Pick the new individual from the new generation if better than previous, else promote the old individual to the next generation unchanged
Then two measures are taken to evaluate the population
- Tracking Error (Fitness, smaller the better): Tracking error is defined as the root-mean-square of differences between the log returns of the index and the individual.
- Diversity (smaller the better): Diversity is defined as the average standard deviation of each chromosome across all individuals. This is an indication that the individuals in this population are similar or not, as we get closer to the optimal solution the variations will become less, as all individuals will become closer to that optimal.
Also I’ve chosen to use Excel as a front end to make use of its charting and spread sheet functions. Excel add-ins, if not carefully designed, may lead to making Excel non-responsive, and one of the goals was to keep excel responsive to enable saving, and cancellation. This is why I split the add-in in two parts. The one that coupled and driven by excel creates a concurrency Agent that will carry on the calculations. Then an integer handle to that agent is created and returned to Excel. Every time Excel wishes to receive the results, or check completion it communicates with the first part with that handle. This allowed decoupling of background processing and UI, while allowing good communication between both.
Due to the fact that each generation depends on the previous one, the loop that evolves the generations can’t be parallelized, but the evolution process itself is parallelizable, as individuals are free to evolve independent of each other. The first chance for parallelizing this application was inside the Evolve method, that had a for loop that executes mutation, recombination, and selection for each individual. Profiling the serial version with the sampling profiler, we find out It contributes to 86% of the total work done by the application. Each loop has read access to all elements of the old generation, and only write-access to individual (i) of the new generation during selection. So, there is no sharing contention, by simply changing the serial for loop to parallel_for the application started to scale up. This sample also show how easy it is to use C++0x lambdas to make this replacement close to a mirror image.
1: if( _asParallel )
2: {
3: parallel_for( 0, _numIndividuals, [&]( size_t i )
4: {
5: Individual y = Mutate( p, i );
6:
7: Recombine( p, i, y );
8: Repair( y );
9: Select( newPopulation, i, y );
10: } );
11: }
12: else
13: {
14: for( size_t i = 0 ; i < _numIndividuals ; ++i )
15: {
16: Individual y = Mutate( p, i );
17:
18: Recombine( p, i, y );
19: Repair( y );
20: Select( newPopulation, i, y );
21: }
22: }
Doing some measurements against the serial implementation, the scaling effecieny[1] was less than 70%. So I executed the following to debug performance:
· started to use the Concurrency Visualizer that comes with Visual Studio 2010.
· turned on tracing by calling (Concurrency::EnableTracing()[2];). This adds markers to my profiler traces to detect where my parallel loops started and ended.
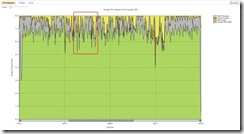
In the "CPU Utilization" view[3] it was clear that the CPUs are not all completely utilized all the time. So I zoomed in at one interesting point and switched to the "Threads" view[4] to have a better look at what the threads are doing.
Figure 2: CPU Utilization view, in Concurrency Visualizer
Looking at the thread view I started to understand why I was unable to utilize all the CPUs. My population was on the order of 200 individuals and the work associated with each was a few thousands cycles. Each core of my 8 cores was processing 25 individuals. Due to randomization not all of cores completed their work at the same time. Near the end, when there was not enough work to keep all CPUs busy, some cores were left without work to do while other cores completed the processing for the parallel loop. In the sample image you can see that only 3 threads are executing when we have 5 idle CPUs. This suggested that if we added a second smaller size work, those additional tasks could get executed during the idle times of the waiting threads. Another interesting point was a serial part between the iterations contributed to 25% of execution of each Evolution loop.
Figure 3: Threads view in Concurrency Visualizer
Now that we know what was blocking the performance let us see how we fixed each one.
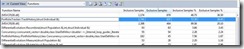
First the fine grain concurrency: we need to find small work that is executed frequently and try to parallelize it. Going back to the sampling profiler, I found that the error tracking function simply calculates the log of the difference between an individual and the original index. It executed 72% of the time, and the exclusive work[5] was 26%. We call it during selection, reparation, creation of new population, and in reporting the best individual.
Figure 4: Functions view from the Sampling Profiler
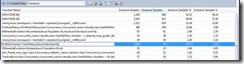
Again, I’ve replaced the for loop with parallel_for. But this is not safe as the result vector needs protection. The Concurrency Runtime provides a cooperative reader writer lock, and even better a concurrent vector that is thread-safe. Concurrent_vector and is compatible with std::vector except for deleting elements so the change was as simple as changing from std::vector<> to Concurrency::concurrent_vector<>. After completing the changes the code again looked identical, involving no major changes. This reduced the exclusive work for that function to less than 1.2%.
1: PortfolioHistory PortfolioTracker::TrackHistory( Individual &w )
2: {
3: Concurrency::concurrent_vector<
4: double,
5: UserDefAlloc< double > > prices;
6:
7: if( _asParallel )
8: {
9: parallel_for( 0, pricesSize, [&]( size_t j )
10: {
11: double newPrice = 0.0;
12: for( size_t i = 0 ; i < indexes.size() ; ++i )
13: {
14: int index = indexes[ i ];
15: newPrice += _data._Members[ index ]._Prices[ j ]
16: * w._Chromosomes[ index ];
17: }
18: prices[ j ] = newPrice;
19: } );
20: }
21: else
22: {
23: for( size_t j = 0 ; j < pricesSize ; ++j )
24: {
25: double newPrice = 0.0;
26: for( size_t i = 0 ; i < indexes.size() ; ++i )
27: {
28: int index = indexes[ i ];
29: newPrice += _data._Members[ index ]._Prices[ j ]
30: * w._Chromosomes[ index ];
31: }
32: prices[ j ] = newPrice;
33: }
34: }
35: }
36:
37: template<typename T>
38: class UserDefAlloc
39: {
40: public:
41: pointer allocate(size_type count, const void *)
42: {
43: return (allocate(count));
44: }
45:
46: pointer allocate(size_type count)
47: {
48: // Call ConcRT Sub Allocator's Alloc
49: pointer ptr = static_cast<pointer>(Alloc(count * sizeof(T)));
50: return ptr;
51: }
52:
53: void deallocate(pointer ptr, size_type)
54: {
55: // Call ConcRT Sub Allocator's Free
56: Free(ptr);
57: }
58:
59: void construct(pointer ptr, const T& val)
60: {
61: // Call the std construct to construct obj@ptr with value val
62: std::_Construct(ptr, val);
63: }
64:
65: void destroy(pointer ptr)
66: {
67: // Call the std destroy to call destructor obj@ptr
68: std::_Destroy(ptr);
69: }
70: };
Figure 5: Functions view from the sampling profiler
Second, the non parallelizable part. This part starts once the evolution ends. We need to decide which generation to pick, the new one or previous ones comparing the best individual from both. then we need to format the result and cache it, so Excel can query the results later on. The serial implementation execute this in serial and we need a reader writer lock to protect the results list until the results are updated with the new one while the parallel version executes the result preparation in parallel using a background task group, then asynchronously sends the results to an unbounded buffer. The second part of the add-in that is coupled with Excel, will use this unbounded buffer to retrieve, and display the results.
1: TG.run( [&]()
2: {
3: //
4: // do the exhaustive copy in the back ground
5: //
6: latestResult->_Population = *best;
7: } );
8:
9: TG.run( [&]()
10: {
11: //
12: // calculate the track history of the best individual
13: //
14: if ( best->_Optimum != Population::uninitialized )
15: {
16: latestResult->_PopulationBest =
17: TrackHistory( best->_Individuals[ best->_Optimum ] );
18: }
19: else
20: {
21: latestResult->_PopulationBest =
22: PortfolioHistory(
23: MyVector( double )( _data._Index._Prices.size(), 0.0 ) );
24: }
25: } );
26:
27: //
28: // execute other code,
29: // then at the end of the function, wait for the task_group
30: //
31:
32: TG.wait();
33:
34: if( _latestResultQueue != NULL )
35: {
36: //
37: // asynchronously send the result to Excel
38: //
39: asend<PortfolioTrackerProgressEventArgs *>(
40: _latestResultQueue,
41: latestResult );
42: }
After fixing these two parts, the program speedup is 3.5x on 4 core, and On 8 core speedup is 6x.
To summarize:
The Concurrency Runtime enables developers to parallelize existing serial applications. Visual Studio 2010 is packed with features that enable the developer to debug, profile, all integrated in one tool.
· The Concurrency Runtime features used
- Agents to work in the background and drive the calculation. While Excel is still responsive.
- Message blocks to pipe line the results to Excel
- Parallel For to replace some for loops
- Task Groups to execute background tasks and wait on them
- Alloc to improve / remove memory contention
- Concurrent vector to remove sharing locks
· Visual Studio 2010 features used
Links and Blogs:
· ConcRT:
- https://msdn.microsoft.com/en-us/library/dd504870(VS.100).aspx
- https://blogs.msdn.com/nativeconcurrency/
- Samples: https://code.msdn.microsoft.com/concrtextras
· Profiler:
- https://msdn.microsoft.com/en-us/library/ee624188(VS.100).aspx
- https://blogs.msdn.com/visualizeparallel/
- https://blogs.msdn.com/hshafi/
Videos:
· ConcRT:
- https://channel9.msdn.com/shows/Going+Deep/Parallel-Computing-in-Native-Code-New-Trends-and-Old-Friends/
- https://channel9.msdn.com/posts/Charles/Parallel-Computing-Platform-Asynchronous-Agents-for-Native-Code/
- https://channel9.msdn.com/posts/Charles/The-Concurrency-Runtime-Fine-Grained-Parallelism-for-C/
- https://channel9.msdn.com/posts/philpenn/concrtppl/
· Profiler:
- https://channel9.msdn.com/posts/philpenn/Concurrent-Visualization-Techniques-in-the-VS2010-Profiler/
- https://channel9.msdn.com/shows/Going+Deep/Visualizing-Concurrency-Inside-the-Concurrency-Visualizer-Profiling-Tool/
- https://channel9.msdn.com/posts/philpenn/New-Parallel-Capabilities-of-the-Visual-Studio-2010-Profiler/
- https://channel9.msdn.com/posts/philpenn/Thread-Visualization-in-the-VS2010-Profiler/
- https://channel9.msdn.com/posts/philpenn/Thread-Blocking-Analysis-in-the-VS2010-Profiler/
References
- [1] D.K. Tasoulis, N.G. Pavlidis, V.P. Plagianakos & M.N. Vrahatis: Parallel Differential Evolution
- [2] M.G. Epitropakis, V.P. Plagianakos, & M.N. Vrahatis: Non-Monotone Differential Evolution
[1] It is the speed up divided by cores, so if the parallel implementation is 4x faster on 8 core then the scalability is 50%.
[2] Enabling tracing will show markers for parallel_for, parallel_foreach, and parallel_invoke. Whether they are called directly or nested inside each other. The marker will mark span from the first iteration to the last executed one.
[3] https://msdn.microsoft.com/en-us/library/dd627195(VS.100).aspx
[4] https://msdn.microsoft.com/en-us/library/dd627193(VS.100).aspx
[5] Work done exclusively inside that function, not inside the functions it calls into.