A KMeans example for Spark MLlib on HDInsight
Today we will take a look at Sparks's module for MLlib or its built-in machine learning library Sparks MLlib Guide . KMeans is a popular clustering method. Clustering methods are used when there is no class to be predicted but instances are divided into groups or clusters. The clusters hopefully will represent some mechanism at play that draws the instance to a particular cluster. The instances assigned to the cluster should have a strong resemblance to each other. A typical use case for KMeans is segmentation of data. For example suppose you are studying heart disease and you have a theory that individuals with heart disease are overweight. You have collected data from individuals with and without heart disease and measurements of their weight like their body mass index, waist-to-hip ratio, skinfold thickness, and actual weight. KMeans is used to cluster the data into groups for further analysis and to test the theory. You can find out more about KMeans on Wikipedia Wikipedia KMeans .
The data that we are going to use in today's example is stock market data with the ConnorsRSI indicator. You can learn more about ConnorsRSI at ConnorsRSI. Below is a sample of the data. ConnorsRSI is a composite indicator made up from RSI_CLOSE_3, PERCENT_RANK_100, and RSI_STREAK_2. We will use these attributes as well as the actual ConnorsRSI (CRSI) and RSI2 to pass into our KMeans algorithm. The calculation of this data is already normalized from 0 to 100. The other columns like ID, LABEL, RTN5, FIVE_DAY_GL, and CLOSE we will use to do further analysis once we cluster the instances. They will not be passed into the KMeans algorithm.
Sample Data (CSV): 1988 instances of SPY SPY
| Column | Description |
| ID | Used to uniquely identify the instance. date:symbol |
| Label | If the close was up or down from the previous days close. |
| RTN5 | The return from the past 5 days. |
| FIVE_DAY_GL | The return from the next 5 days. |
| Close | Closing price. |
| RSI2 | Relative Strength Index (2 days). |
| RSI_CLOSE_3 | Relative Strength Index (3 days). |
| RSI_STREAK_2 | Relative Strength Index (2 days) for streak durations based on the closing price. |
| PERCENT_RANK_100 | The percentage rank value over the last 100 days. This is a rank that compares todays return to the last 100 returns. |
| CRSI | The ConnorsRSI Indicator. (RSI_CLOSE_3 + RSI_STREAK_2 + PERCENT_RANK) / 3 |
| ID | LABEL | RTN5 | FIVE_DAY_GL | CLOSE | RSI2 | RSI_CLOSE_3 | PERCENT_RANK_100 | RSI_STREAK_2 | CRSI |
| 2015-09-16:SPY | UP | 2.76708 | -3.28704 | 200.18 | 91.5775 | 81.572 | 84 | 73.2035 | 79.5918 |
| 2015-09-15:SPY | UP | 0.521704 | -2.29265 | 198.46 | 83.4467 | 72.9477 | 92 | 60.6273 | 75.1917 |
| 2015-09-14:SPY | DN | 1.77579 | 0.22958 | 196.01 | 47.0239 | 51.3076 | 31 | 25.807 | 36.0382 |
| 2015-09-11:SPY | UP | 0.60854 | -0.65569 | 196.74 | 69.9559 | 61.0005 | 76 | 76.643 | 71.2145 |
| 2015-09-10:SPY | UP | 0.225168 | 1.98111 | 195.85 | 57.2462 | 53.9258 | 79 | 65.2266 | 66.0508 |
| 2015-09-09:SPY | DN | 1.5748 | 2.76708 | 194.79 | 42.8488 | 46.1728 | 7 | 31.9797 | 28.3842 |
| 2015-09-08:SPY | UP | -0.12141 | 0.521704 | 197.43 | 73.7949 | 64.0751 | 98 | 61.2696 | 74.4483 |
| 2015-09-04:SPY | DN | -3.35709 | 1.77579 | 192.59 | 22.4626 | 31.7166 | 6 | 28.549 | 22.0886 |
The KMeans algorithm needs to be told how many clusters (K) the instances should be grouped into. For our example let's start with two clusters to see if they have a relationship to the label, "UP" or "DN". The Apache Spark scala documentation has the details on all the methods for KMeans and KMeansModel at KMeansModel
Below is the scala code which you can run in a zeppelin notebook or spark-shell on your HDInsight cluster with Spark. HDInsight
import org.apache.spark.mllib.linalg.Vectors
import org.apache.spark.mllib.clustering.KMeans
import org.apache.spark.sql.functions._
// load file and remove header
val data = sc.textFile("wasb:///data/spykmeans.csv")
val header = data.first
val rows = data.filter(l => l != header)
// define case class
case class CC1(ID: String, LABEL: String, RTN5: Double, FIVE_DAY_GL: Double, CLOSE: Double, RSI2: Double, RSI_CLOSE_3: Double, PERCENT_RANK_100: Double, RSI_STREAK_2: Double, CRSI: Double)
// comma separator split
val allSplit = rows.map(line => line.split(","))
// map parts to case class
val allData = allSplit.map( p => CC1( p(0).toString, p(1).toString, p(2).trim.toDouble, p(3).trim.toDouble, p(4).trim.toDouble, p(5).trim.toDouble, p(6).trim.toDouble, p(7).trim.toDouble, p(8).trim.toDouble, p(9).trim.toDouble))
// convert rdd to dataframe
val allDF = allData.toDF()
// convert back to rdd and cache the data
val rowsRDD = allDF.rdd.map(r => (r.getString(0), r.getString(1), r.getDouble(2), r.getDouble(3), r.getDouble(4), r.getDouble(5), r.getDouble(6), r.getDouble(7), r.getDouble(8), r.getDouble(9) ))
rowsRDD.cache()
// convert data to RDD which will be passed to KMeans and cache the data. We are passing in RSI2, RSI_CLOSE_3, PERCENT_RANK_100, RSI_STREAK_2 and CRSI to KMeans. These are the attributes we want to use to assign the instance to a cluster
val vectors = allDF.rdd.map(r => Vectors.dense( r.getDouble(5), r.getDouble(6), r.getDouble(7), r.getDouble(8), r.getDouble(9) ))
vectors.cache()
//KMeans model with 2 clusters and 20 iterations
val kMeansModel = KMeans.train(vectors, 2, 20)
//Print the center of each cluster
kMeansModel.clusterCenters.foreach(println)
// Get the prediction from the model with the ID so we can link them back to other information
val predictions = rowsRDD.map{r => (r._1, kMeansModel.predict(Vectors.dense(r._6, r._7, r._8, r._9, r._10) ))}
// convert the rdd to a dataframe
val predDF = predictions.toDF("ID", "CLUSTER")
The code imports some methods for Vector, KMeans and SQL that we need. It then loads the .csv file from disk and removes the header that have our column descriptions. We then define a case class, split the columns by comma and map the data into the case class. We then convert the RDD into a dataframe. Next we map the dataframe back to an RDD and cache the data. We then create an RDD for the 5 columns we want to pass to the KMeans algorithm and cache the data. We want the RDD cached because KMeans is a very iterative algorithm. The caching helps speed up performance. We then create the kMeansModel passing in the vector RDD that has our attributes and specifying we want two clusters and 20 iterations. We then print out the centers for all the clusters. Now that the model is created, we get our predictions for the clusters with an ID so that we can uniquely identify each instance with the cluster it was assigned to. We then convert this back to a dataframe to analyze.
Below is a subset of the allDF dataframe with our data.
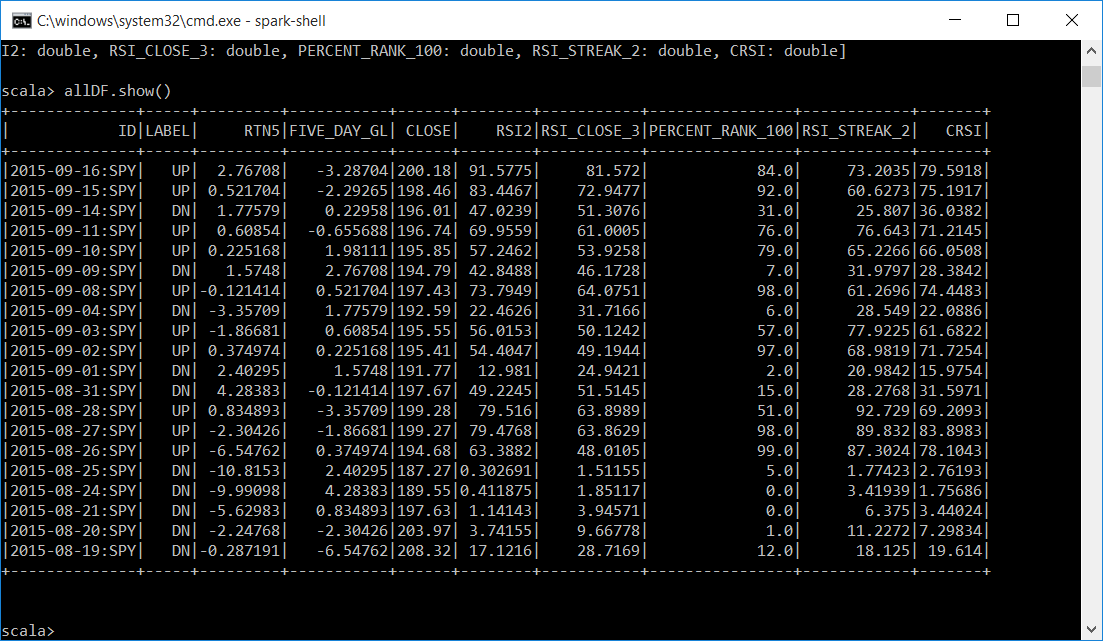
Below is a subset of our predDF dataframe with the ID and the CLUSTER. We now have a unique identifier and which cluster the KMeans algoritm assigned it to. Also displayed is the mean for each of the attributes passed into the KMeans algorithm for each cluster. Cluster 0 and Cluster 1. You can see that the means are very close in each cluster. For Cluster 0 it is around 27 and for cluster 1 it is around 71.
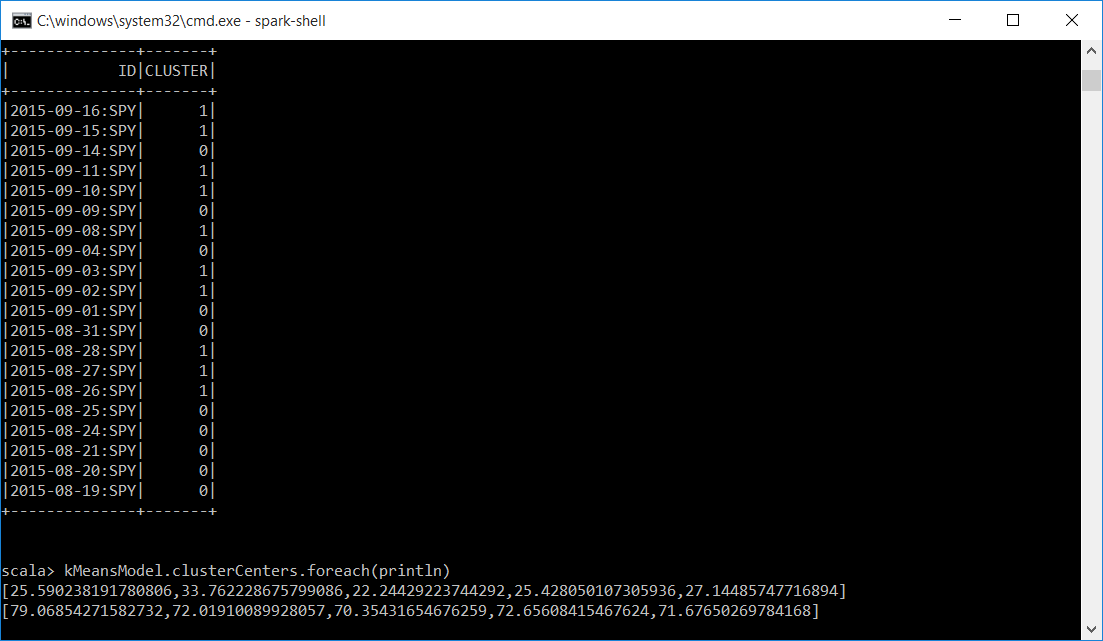
Because the allDF and predDF dataframes have a common column we can join them and do more analysis.
// join the dataframes on ID (spark 1.4.1)
val t = allDF.join(prdDF, "ID")
Now we have all of our data combined with the CLUSTER that the KMeans algorithm assigned each instance to and we can continue our investigation.
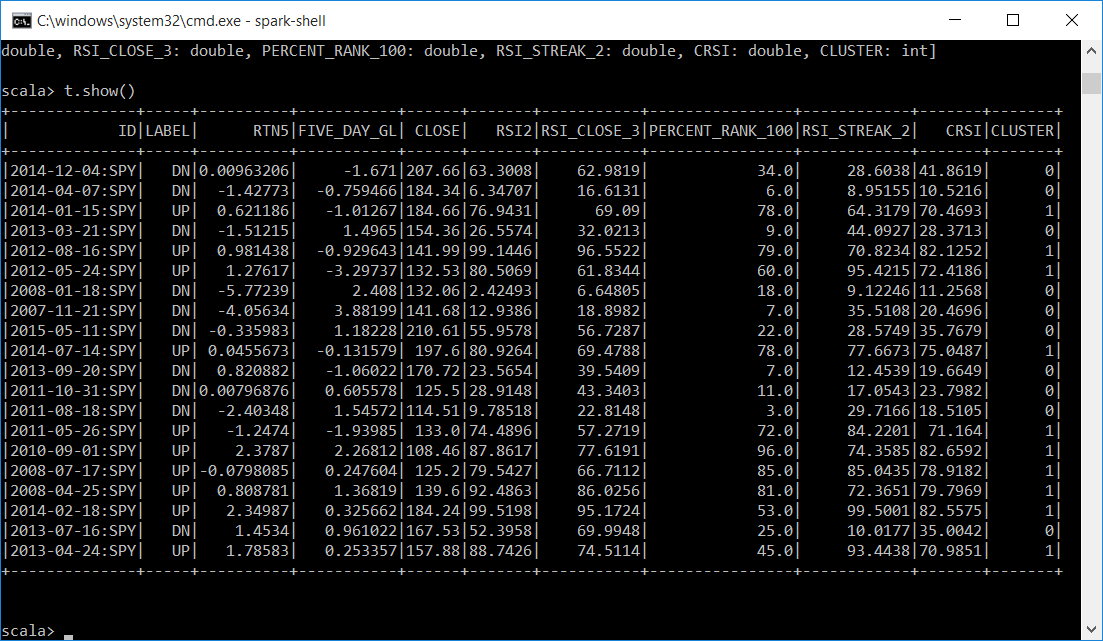
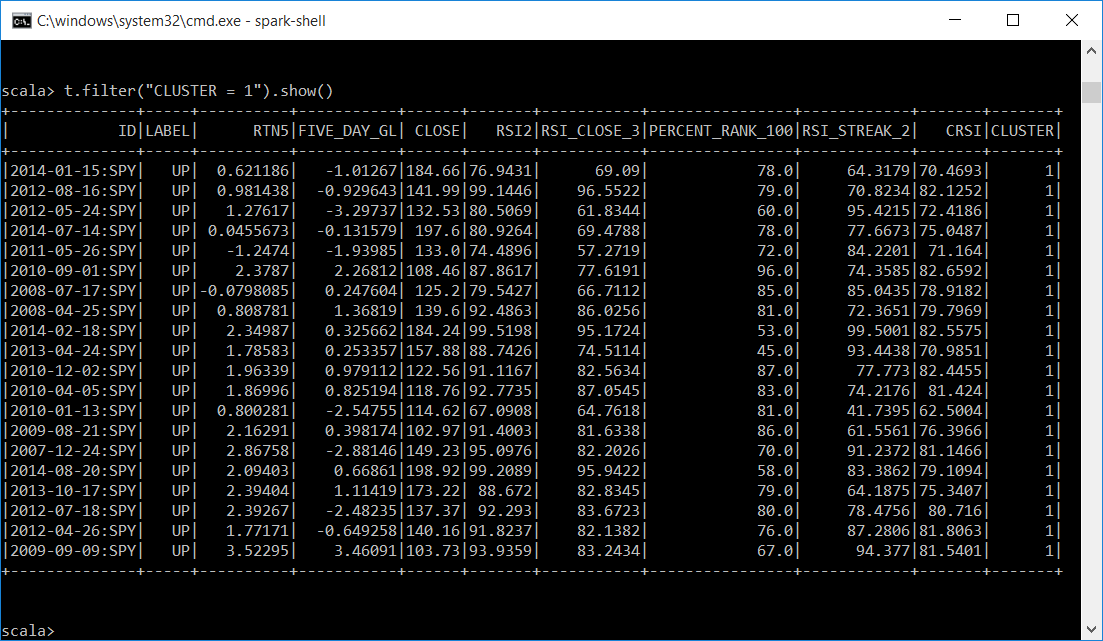
Let's display a subset of each cluster. It looks like cluster 0 is mostly DN labels and has attributes averaging around 27 like the centers of the clusters indicated. Cluster 1 is mostly UP labels and the attributes average is around 71.
// review a subset of each cluster
t.filter("CLUSTER = 0").show()
t.filter("CLUSTER = 1").show()
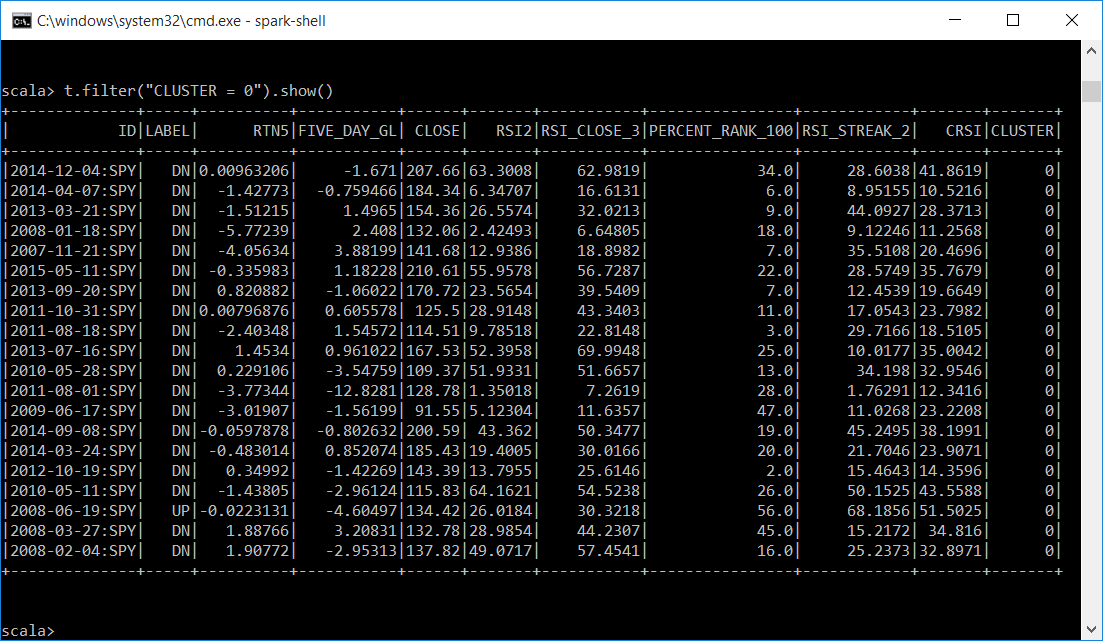
Let's get descriptive statistics on each of our clusters. This is for all the instances in each cluster and not just a subset. This gives us the count, mean, stddev, min, max for all numeric values in the dataframe. We filter each by CLUSTER.
// get descriptive statistics for each cluster
t.filter("CLUSTER = 0").describe().show()
t.filter("CLUSTER = 1").describe().show()
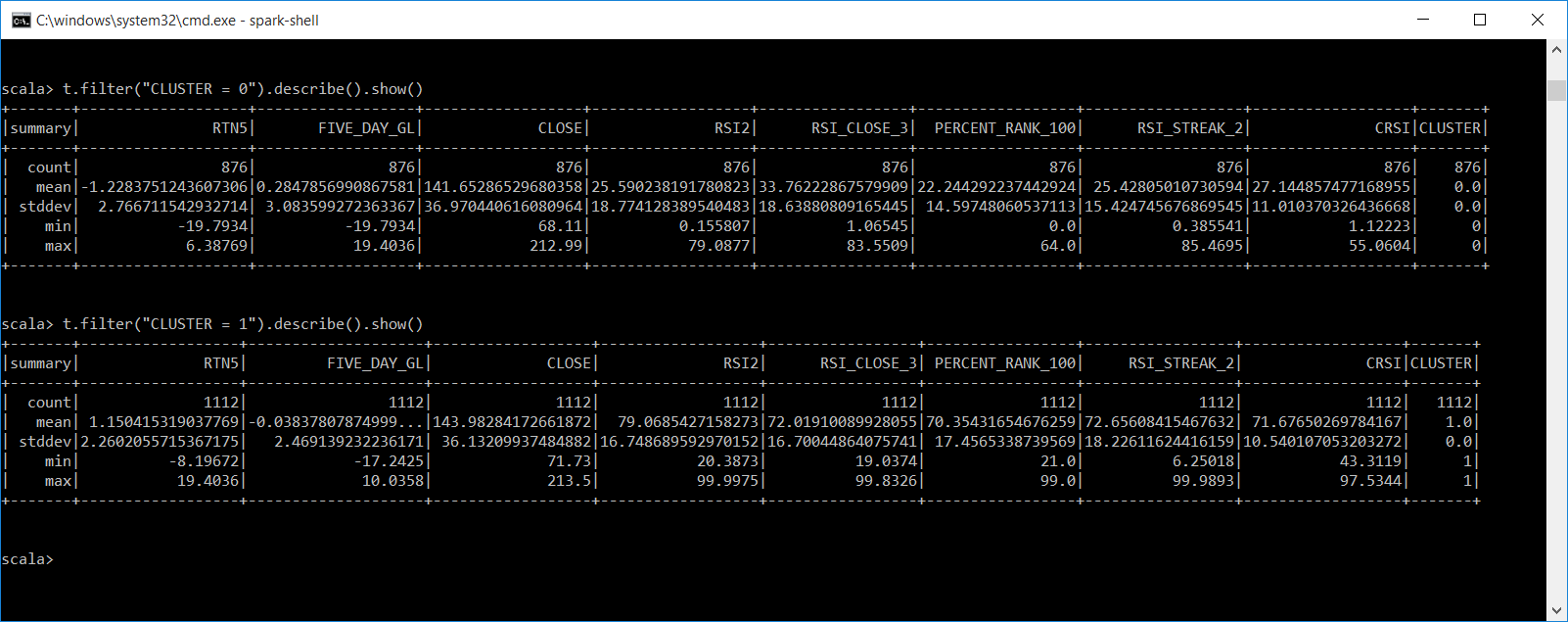
So what can we infer from the output of our KMeans clusters?
- Cluster 0 has lower ConnorsRSI (CRSI), with a mean of 27. Cluster 1 has higher CRSI, with a mean of 71. Could these be areas to initiate buy and sells signals?
- Cluster 0 has mostly DN labels, and Cluster 1 has mostly UP labels.
- Cluster 0 has a mean of .28 % gain five days later, while cluster 1 has a loss of .03 five days later.
- Cluster 0 has a mean loss of 1.22% five days before and cluster 1 has a gain of 1.15% five days before. Does this suggest markets revert to their mean?
- Both clusters have min\max of 5 day returns between positive 19.40% to a loss of 19.79%.
This is just the tip of the iceberg with further questions, but gives an example of using HDInsight and spark to start your own KMeans analysis. Spark MLlib has many algorithms to explore including SVMs, logistic regression, linear regression, naïve bayes, decision trees, random forests, basic statistics, and more. The implementation of these algorithms in spark MLlib is for distributed clusters so you can do machine learning on big data. Next I think I'll run the analysis on all data for AMEX, NASDAQ and NYSE stock exchanges and see if the pattern holds up!

Bill