Research Life Science Projects on the Windows Azure Cloud (Part 2)
Life sciences has been one of the most productive areas of research using cloud computing. There are many reasons for this. First and foremost is the fact that the genomics revolution is now driven by large data and many important discoveries are being made by exploiting large scale data analytics and machine learning. Second, bioinformatics is increasingly becoming a discipline that requires sharing access to important data collection over the Internet. The massive data centers that form the foundation of cloud computing were originally designed to support large scale data analytics for Internet search engines and for delivering data to thousands of users concurrently. These two capabilities fit nicely with the needs of bioinformatics.
Big Compute and Bioinformatics
In the case of using the cloud for large scale data analytics we have already mentioned the work of Wuchun Feng of Virginia Tech (see part 1 of this series). However there are many other examples. One very impressive example comes from the work of my colleagues at Microsoft Research in David Heckerman’s eScience team. They used Windows Azure to conduct a genome-wide association study using 27,000 computing cores for 72 hours. They were looking for the genetic markers associated with a number of important human diseases. One of the big challenges in such a large study is to statistically correct for the “confounding” caused by the relatedness of individuals in the subject pool. Traditional methods to do this don’t scale well, so they designed a new linear time algorithm called a Factored Spectrally Transformed Linear Mixed Model (FaST-LMM). Using this they were able to analyze 63 billion pairs of genetic markers. Finally they put the results into the Azure Marketplace where users can validate results from their own studies. To learn more read this article.
Another examples comes from Nikolas Sgourakis. When he was a postdoc at the University of Washington in the Baker lab he undertook a project “Protein Folding with Rosetta@home in the Cloud”. The scientific challenge was to elucidate the structure of a molecular machine called the needle complex, which is involved in the transfer between cells of dangerous bacteria, such as salmonella and e-coli. This was accomplished by porting Rossetta to Windows Azure using 2,000 cores. Results of this work can be found in this paper in nature (full article here)
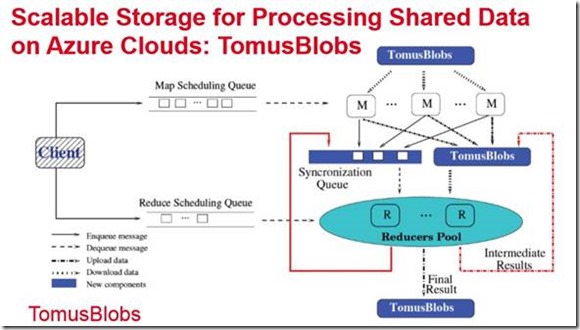
Gabrial Antoniu and Bertrand Thirion from the famous French research lab INRIA led the project “A-Brain: Large-scale Joint Genetic and Neuroimaging Data Analysis on Azure Clouds”. It is believed that several brain diseases have genetic origin or the severity of the disease is related to genetic factors. Consequently it may be possible to use a patient’s genetic profile to predict the response of to a given treatment. With help from my colleagues at the Microsoft Advanced Technology Lab in Europe they were trying to find the correlation between certain brain markers and genetic data. By using neuroimaging and genetic SNP data from a thousand patients they built an analysis pipeline in the cloud using MapReduce-style computation. Using a ridge regression approach the team has provided the first statistical evidence of the heritability of function signals in basal ganglia related to a failed stop task (a standard cognition metric). One of the major challenges they faced was the way standard MapReduce computations are done. As is often the case when MapReduce is in the inner loop of an iterative computation the storage updates and locality are a bottleneck. To solve this problem they introduced something they called TomusBlobs which is a storage system based on aggregated virtual disks and versioning to provide lock-free concurrent writes. They were then able to scale up to 1000 cores on 3 Azure data centers and a performance improvement of 50% over standard Azure Blob storage.
A summary of their work is available in this presentation.
Two other projects used Windows Azure for large scale bioinformatics work. Michele Di Cosmo and Angela Sanger of the University of Trento Centre for Computational and Systems have developed BetaSIMm on Windows Azure. BetaSIMm is a dry experiment simulator, driven by BlenX—a stochastic, process algebra-based programming language for modeling and simulating biological systems as well as other complex dynamic systems. The second is from Theodore Dalamagas, Thanassis Vergoulis, and Michalis Alexakis of Athena RC. They ran a project “Targets on the Cloud: a Cloud-Based MicroRNA Target Prediction Platform”. In this effort they set up target prediction methods on the Windows Azure platform to provide real-time target prediction services. Our aim is to meet the requirements for efficient calculation of targets and efficient handling of the frequent updates in the miRNA and gene databases.
The Cloud for Providing Bioinformatics Services over the Internet.
The Trento BetaSiMm on Azure project and the Athena projects mentioned above were both part of an EU funded collaboration called Venus-C with cloud resources provided by Microsoft. Two other Venus-C projects looked at using the cloud as a delivery mechanism for bioinformatics tools over the Internet. The importance of providing tools for the community cannot be underestimated. The vast majority of scientific researchers do not want to develop, install and debug cloud application. They want to get on with their science. The open source science community has built a large collection of tools that are used by thousands of researchers. Most researchers download and install these tools on local resources, but this is often a suboptimal approach because the servers upon which they run need to be maintained, the software updates have to be tracked and installed and much of the data they need lies in large institutional repositories. A better solution is for a community of researchers to support standard collections of services and data as a cloud service that can be access by anybody over the web.
There are two approaches for using the cloud as the host for research tools. The first is to provision the tool collections as web services. This is the approach taken by the collaboration that grew out of the Venus-C project and later funded directly by Microsoft Research. Ignacio Blanquer,Abel Carrión and Vicente Hernández of the Universidad Politécnica de Valencia and Paul Watson and Jacek Cala from the University of Newcastle and the Trust-IT group teamed up to produce a prototype community bioinformatics service called cloud4science. What they have done is prototyped and developed a client tool that provides the same interface as the conventional bioinformatics tools (such as BLAST, bwa, fasta, bowtie, BLAT, and ssaha) that are used in local computers, but linking them to a powerful processing service on the cloud. The initial target is next generation sequencing challenges. A secondary goal is to demonstrate a model for self-sustainability of such services by the user community.
The system is built as a configurable pipeline for the detection of mutations. Currently it allows the users to define various workflows for sequence trimming, quality analysis, mapping, variant discovery and visualization. Users have their own remote storage space on the cloud and access to preloaded reference data (FASTA Genomes of HG19 and Drosophila Melanogaster + Bowtie2 indexes). The
· Basic pe-procressing of FASTQ/FASTA files through SEQTK.
· Quality analysis of sequence files with FASTQC.
· Alignment and BAM generation using BWT (Bowtie2)
· Variant discovery with GATK.
· Visualization of BAM and VCF files with GenomeMaps.
Integrated visualization uses bandwidth-friendly tools so that data can be explored without downloading.
The current version is still a prototype and we are looking for community input on how it can be improved.
Recent publications from this group include
o Ignacio Blanquer, Goetz Brasche, Jacek Cala, Fabrizio Gagliardi, Dennis Gannon, Hugo Hiden, Hakan Soncu, Kenji Takeda, Andrés Tomás, Simon Woodman, Supporting NGS pipelines in the cloud, 2013 - journal.embnet.org
o D Lezzi, R Rafanell, A Carrión, Enabling e-Science applications on the Cloud with COMPSs, Euro-Par, 2012 – Springer
o J Cała, H Hiden, S Woodman, P Watson, Fast Exploration of the QSAR Model Space with e-‐Science Central and Windows Azure. 2012 - esciencecentral.co.uk
Further details about this project can be found in Ignacio’s presentation at the BIO-IT Clinical Genomics & Informatics conference in Lisbon this week.
It should be noted that these are only two of many examples and we will describe more of them later.
I mentioned above that there were two approaches to providing cloud science services over the Internet. The ideal solution is the website service such as the Blanquer/Watson example. There is a downside to this approach that make it unappealing to some researchers. Specifically those researchers who would like a more customized view that they can control and yet still have access to the common data collections. To make it easy to serve this community it is possible to populate an entire virtual machine with all the basic tools and then make this VM image available on-line so that it can be deployed by anybody. This can save the developer a great deal of time and effort because many of the deployment tasks are already done. The user can then simply weave together the needed components with a script of their own design. The Microsoft Open Technologies VMDepot has been set up to provide a collection of such VMs. One of the standard tools for bioinformatics is Galaxy and it is available as a VM image from the Depot as part of the biolinux platform. In a later post to this blog we will describe the contents and use of the preconfigured VM in greater detail.